How are ASF viruses classified?
The African swine fever virus (ASFV) has been classified into various genotypes based on the sequence of the B646L gene, which encodes the p72 protein, the major component of the virus. Based on this sequence, up to 24 different genotypes have been described as circulating around the African continent.
Outside Africa, only the following genotypes have been declared as being in circulation:

- Genotype I: currently circulating in Asia, and responsible for the pandemic that appeared in the 1960s.
- Genotype II: the predominant genotype in the current pandemic, which started in Georgia in 2007.
The European reference laboratory proposed the classification by genetic groups within genotype II based on differences in the sequences of six variable regions of the genome of the virus.
| CVR | IGR I73R/I329L | O174L |
| K145R | IGR MGF505 9R/10R | IGR I329L-I215L |
Based on the differences found in these regions, a total of 28 genetic groups were described, with genetic group #1 represented by the Georgia 2007 strain.
The proposed nomenclature for the strain found in Barcelona is SP25WB2611, abbreviated to Sp25.
The strain found in Barcelona (Sp25) is different from those known until now and has therefore been classified in a new genetic group: #29.
Some genetic groups have only been isolated in domestic pigs, others only in wild boars, and others in both animals. Some genetic groups, such as #3 or #19, are found in different countries, but most have only been detected in a specific geographical area. There are also areas with more than one genetic group in circulation, e.g. the Russian Federation (#1, #2, #3, #4, and #7) and Romania (#3, #7, #19, #21, #22, and #24).
The genome of the Sp25 strain has a large deletion of approximately 10Kb at the left end of the genome, together with a total of 27 SNPs (Single Nucleotide Polymorphisms, i.e. point mutations that cause changes of a single nucleotide in the virus sequence) compared to the Georgia 2007 strain.
What is the distribution in terms of space and time of the sequences available in Genebank, with which the new strains are compared?
Although many strains of the ASFV have been isolated in Europe since 2007, the number of sequences available is very low in comparison.
Knowledge of the geographical distribution of genetic groups depends on the number of samples sequenced and published in each country. In many countries in which ASF is present, the results are not sequenced and/or published.
The sequence analyses of the Sp25 strain carried out at the IRB and BSC in Barcelona demonstrate its phylogenetic proximity to strains isolated from outbreaks in European countries after Georgia 2007 spread to the continent (Figure 1). The Sp25 strain appears in a lineage that diverges very early, and it is therefore very different from those causing the most recent outbreaks in central and western Europe. As the tree diagram shows, this sequence is more closely related to viruses detected in Russia in 2019 than to strains isolated in central and western Europe; however, it is still very different from any strain for which the complete genome has been obtained, including those isolated in Russia, differing in the aforementioned deletion and in more than 20 SNPs.
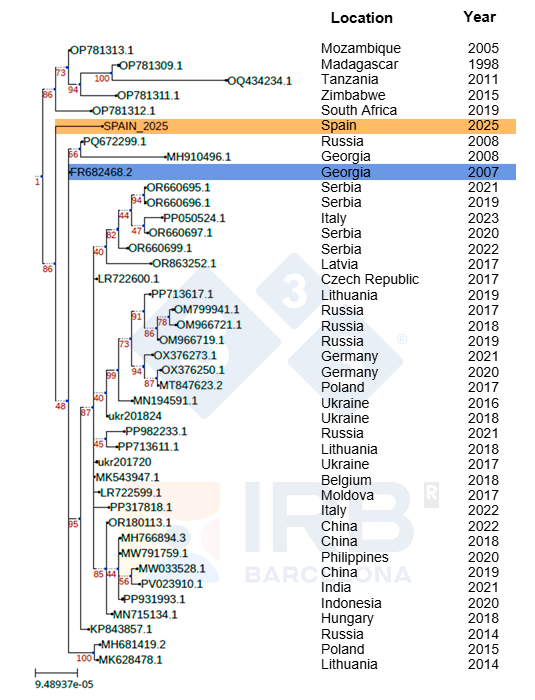

The ASFV is said to be a very large and very stable DNA virus. How many changes must an ASF isolate have for it to be considered a different strain?
Within the context of the ASFV in Europe, the European Reference Laboratory considers that a viral isolate constitutes a different strain from the rest when it is classified as a new genetic group.
The number of SNPs we detect shows us the temporal distancing of a strain from the 'original' Georgia 2007 strain. The ASFV is a double-stranded DNA virus and, therefore, a very stable virus. Given the low mutation rate of the virus, the various SNPs present in Sp25 must have accumulated over an extended period since its divergence from Georgia 2007. According to the latest estimates, this period could be around 10-15 years of evolutionary divergence (estimate made by Dr Toni Gabaldón, IRB).
If it is such a stable virus, can the deletion of 10,000 base pairs observed in the strain found in the Barcelona outbreak occur naturally?
The natural evolutionary dynamics of the virus itself generate this type of genetic deletion. In genotype II, deletions in the same region with different lengths and gene involvement has been found in both Africa (Ambagala et al., 2023) and Europe (Zani et al., 2018; Torresi et al., 2025).

This deletion is used in some vaccine candidates. Does that mean it cannot occur naturally?
This type of deletion in the ASFV genome occurs naturally and can also be generated in the laboratory to create vaccine candidates. It is worth pointing out that nowadays an exogenous marker is usually included in the virus genome (for technical reasons) during the development protocol of a vaccine candidate in the laboratory so that it can be distinguished from the original strain. As a result, vaccine candidates are easily identifiable through sequencing. The Sp25 strain does not carry any type of exogenous marker. In addition, most vaccine candidates are generated from a small group of laboratory strains that are very similar to each other and to the first isolate of the current outbreak: Georgia 2007.
Is recombination between different strains of ASF a common event?
For these recombinations to occur, a host cell must be simultaneously infected with two different strains. This phenomenon is not as common as with RNA viruses with segmented genomes (as is the case with the influenza virus); however, it does occur with the ASFV. Recombinations between genotypes I and II have been reported in China (Zhao et al., 2023), and the same recombinant virus has spread to other countries (Russia and Vietnam). Logically, the more different strains circulating in the same area, the greater the possibility of such recombinations occurring. The pattern of mutations detected in the Collserola strain is distributed throughout the entire genome, and not in a limited region, making an origin by recombination unlikely.
There is a lot of talk about the strain detected in Barcelona not being very virulent. Is that really the case?
The deletion identified in the Sp25 strain could be associated with lower virulence, as has been demonstrated in other strains with deletions in this region. However, there are examples of very virulent strains with similar deletions, which makes the possibility inconclusive. The virulence of the Sp25 strain can only be assessed when an experimental infection is carried out in pigs or wild boars under controlled conditions.
The only certainty we have right now is that it is a lethal strain in wild boar, and that the observed disease pattern is comparable to that described in other European outbreaks caused by strains classified as highly virulent.
Figure 2 shows a temporal reconstruction of wild boar cases in the current ASF outbreak, bearing in mind that the date the carcass was found does not usually coincide with the date of death of the animal. In each case, a plausible time interval for the death of the wild boar is given, estimated based on the state of degradation of the carcass. Thus, although the exact date of the first deaths cannot be determined with any great precision, the estimated intervals in the case of the first carcasses indicate that the mortality associated with the outbreak began months before the first case was detected, probably around September. The ASFV therefore spread undetected for an initial period, likely at a low but sustained level until it was detected. The initial phase of the outbreak, which progresses slowly, corresponds to the introduction and consolidation of the ASFV in the wild boar population, followed by an epidemic phase in which a change in the slope of the curve is observed. It is of interest to note that the progression of this outbreak does not differ greatly from that observed in recent outbreaks in Europe.
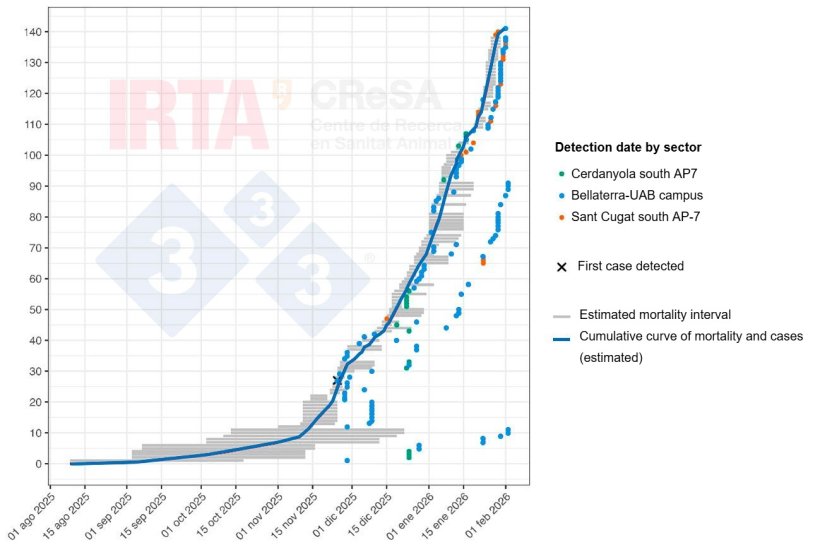
Figure 2. Temporal reconstruction of mortality and cases of ASF in wild boar during the Collserola outbreak. The grey horizontal lines represent the estimated mortality intervals for each animal. The dots show the date on which the carcass or the sick animal was detected. The colour of each dot identifies the geographical sector where the carcass was found (green, Cerdanyola south AP7 sector; blue, Bellaterra-UAB campus sector; and orange, Sant Cugat south AP7 sector). The cross marks the first case detected. The blue line represents the cumulative curve of mortality and cases of ASF.


