Clinical signs
Most infections with M. hyopneumoniae are subclinical. In case of clinical disease, a dry non-productive coughing is the most obvious clinical sign. Under experimental infections, coughing appears from 10 to 14 days post-infection, reaches a maximum at about 4-5 weeks, after which it disappears gradually. Under field conditions, coughing may be present in nursery, grower and finisher pigs and usually a considerable percentage of pigs are affected. In case of concurrent infections with other respiratory pathogens, the clinical signs and lung lesions are more severe. M. hyopneumoniae infections lead to an increase of feed conversion ratio, lower average daily weight gain, and more weight variation between the pigs.

M. hyopneumoniae is also one of the most important pathogens in the porcine respiratory disease complex (PRDC). PRDC includes both bacterial (those potentially involved in enzootic pneumonia) and viral (porcine reproductive and respiratory syndrome virus (PRRSV), porcine circovirus type 2, Aujeszky’s disease virus, swine influenza viruses and porcine respiratory coronavirus) agents. PRDC is clinically characterized by slow growth, decreased feed efficiency, anorexia, fever, cough, and dyspnea in finishing pigs.
Diagnosis
Presumptive diagnosis of mycoplasmal pneumonia is based on the clinical symptoms (chronic non-productive cough with poor performance), typical macroscopic consolidation in the cranioventral parts of the lung (Photo), and the presence of characteristic peribronchiolar and perivascular cuffs and nodules on histopathology. However, neither the clinical symptoms nor the lung lesions are pathognomonic for M. hyopneumoniae. In addition, mixed infections are common resulting in symptoms and lesions that are less typical for M. hyopneumoniae.
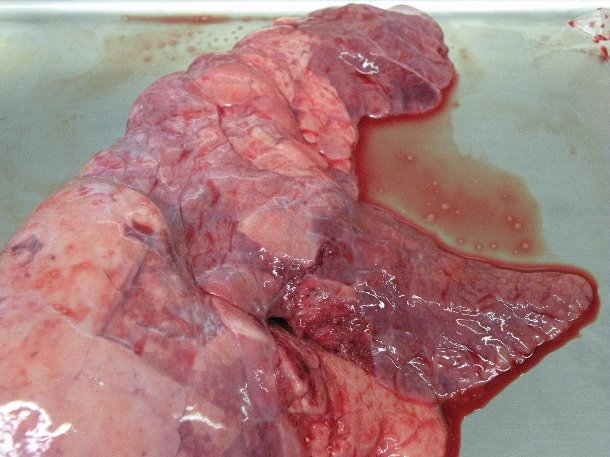
Catarrhal pneumonia in the cranioventral parts of the lung following experimental M. hyopneumoniae infection.
Both clinical symptoms and lesions have been quantified using scoring systems. A coughing index, based on the number of pigs coughing in a given period of time, have been shown to correspond with the infection profile of M. hyopneumoniae under field conditions (Maes et al., 1999; Nathues et al. 2012). Lung lesions can be assessed on dead or necropsied pigs, but also by means of slaughter checks. In order to reliably estimate the prevalence and severity of lung lesions at a herd level, lungs of at least 30 animals, but preferably more, should be inspected (Davies et al., 1995). Slaughter checks are useful to detect subclinical respiratory infections, which can also adversely affect the pig performance. Most scoring methods are based on a visual subjective estimation of the proportion of lung affected surface and/or volume. Therefore, it can involve possible errors. Presence of severe pleurisy may mask pneumonia lesions and healing of lesions during the fattening period may lead to false negative results. Hence, the subjectivity of the scoring method, regression of lung lesions and the non-specificity of these lesions preclude an etiologic diagnosis based solely on examination of gross lung lesions at slaughter.
Isolation of M. hyopneumoniae is still considered as the gold standard for diagnosis, but it is not used in practice because it is very difficult, expensive and time-consuming. Conclusive diagnosis is mostly established by showing (parts of) the pathogen by PCR testing or antigen detection by immunofluorescence, and by using serology. Different PCR tests are available and M. hyopneumoniae can be detected in nasal and tracheal samples, BAL fluid or in lung tissue of dead animals. Nasal swabs can be taken easily in practice from live animals, but the testing is less sensitive than when using tracheal samples or BAL fluid. Vranckx et al. (2012) assessed the infection load in pigs of different age groups in four different herds using qPCR of BAL fluid (Figure 1). The highest numbers were found in the older age groups. These authors also showed a positive correlation in slaughter pigs between the number of M. hyopneumoniae organisms determined by qPCR in BAL fluid and the severity of lung lesions. Taking blood samples of different age groups for detecting serum antibodies allows to investigate a large number of pigs and to have good insight in the infection pattern. Cross-sectional sampling (different age groups during one herd visit) has the advantage that results are obtained quickly. Serial sampling has the advantage that the same group (or pigs) is/are investigated over time and that more information is obtained on the sequence of infections in these groups. Serology for M. hyopneumoniae should be interpreted at group level. There is no correlation between the serum antibody level and the degree of protection using the currently available commercial ELISAs. Seroconversion under field conditions may take 3 to 8 weeks or even more.
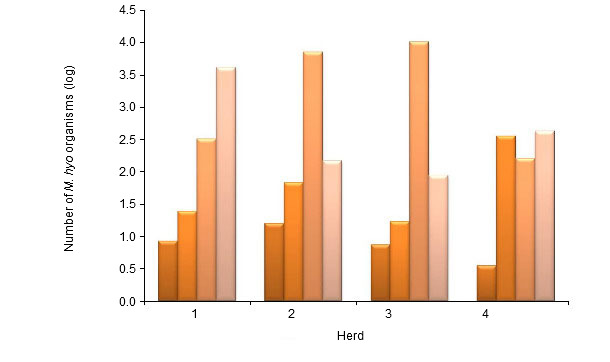
Figure 1. Number of M. hyopneumoniae organisms (mean log number of organisms per ml) tested by nPCR in bronchoalveolar lavage (BAL) fluid in pigs of 6, 10 and 14 weeks and at slaughter age in 4 different herds (Vranckx et al. 2012)