Modelling: a supplementary approach to field and experimental studies
Modelling is more and more frequently used to represent and analyse complex biologic systems with their interactions. George E.P. Box said “essentially all models are wrong, but some are useful” which highlights the main objective of modelling studies: to represent complex systems through a simplification of the system suitable for analysis but capturing essential behaviour of interest and incorporate main processes. Modelling is therefore an important supplementary approach to observational and experimental studies, from which new questions and hypotheses may rise.

Epidemiological studies of risk factors of post-weaning multisystemic wasting syndrome (PMWS) clearly evidenced the dynamics of PCV2 infection in growing pigs as a pivotal event: the earlier the infection, the higher the risk. Deviations in management procedures in the early life of growing pigs were identified as clear risk factors for PMWS and could favour pathogen transmission between pigs. Given the possible vertical transmission of PCV2 and the importance of passive immunity, cross-fostering practices have been suspected to enhance PCV2 transmission from infected litters to susceptible ones. However, the relationships between identified risk factors for PMWS and changes in PCV2 course of infection are not fully understood. By reproducing the complex system of population dynamics subjected to an infectious process, modelling studies can provide useful information about such interactions.
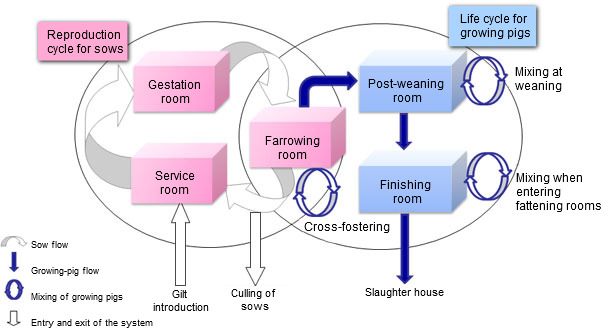
Figure 1. Representation of the pig population dynamic model
Model development
The population dynamics model was built on an individual basis (figure 1). Indeed, the level of representation allowed for featuring the strong relationship between PCV2 infectious statuses of pregnant sows and their progeny and the movements of individual animals, Moreover, this model was made stochastic to account for the biological variability for various individual parameters (number of piglets born per litter, abortion or mortality probabilities...). On the other hand, the epidemiological model was designed on the basis of current knowledge on within-host PCV2 course of infection (figure 2). The main core was a SEIR (Susceptible – Exposed – Infectious – Recovered) model. However, the model had also to account for specific infectious statuses to represent vertical and pseudo-vertical transmissions as well as the impact of maternally derived antibodies. To represent PCV2 course of infection in a realistic way, transmission experiments aiming at characterizing the PCV2 transmission process were carried out. These experiments (picture 1), based on contacts between infected and susceptible animals with sequential monitoring of infected and sentinel piglets, allowed for estimations of the transmission parameter according to different contact structures between individual animals. Moreover, the transmission was found to vary with the time since infection with a transmission peak about 15 days post-infection then declining until day 49 post-infection. Using a similar protocol we also showed a significant reduction of the transmission rate in a vaccinated population. The final model was obtained by merging the population and epidemiological models.
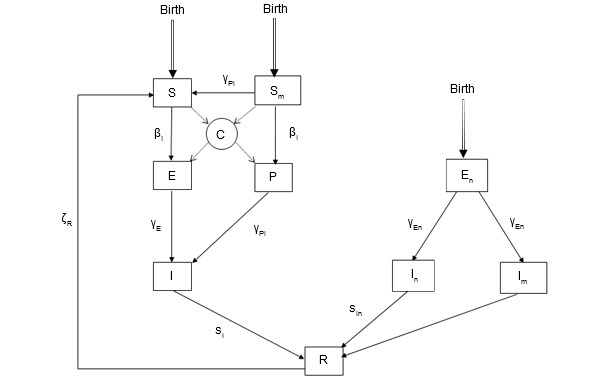
Figure 2. The epidemiological model for PCV2 infection. State variables: S: Susceptible; Sm: Piglets susceptible at birth with passive immunity; E: latency for pigs infected without passive immunity; P: Piglets infected by contact with an infectious individual while having passive immunity; I: Infectious pigs; En: Piglets infected pseudo-vertically by infected semen (latency); In: Infectious piglets (pseudo-vertically infected) without passive immunity; Im: Infectious piglets (pseudo-vertically infected) with passive immunity; R: ‘Recovered’ pigs, this class represents pigs that are no more infectious.

Picture 1: transmission experiment to estimate transmission parameters of the PCV2 model
Lessons learnt from the model
The age at infection, major risk factor for PMWS expression, was studied as the output variable for twelve combinations of management strategies based on cross-fostering practices and growing pigs housing in nursery facilities. Reducing piglets mingling in farrowing and nursery facilities was found to delay the infectious process leading to a lower number of early infections. Clustering piglets by litter in small units after weaning also decreased significantly the probability of early infection. These results showed the possible interactions between identified PMWS risk factors.
Age-specific seroprevalences were recorded for the two extreme management strategies i.e., the most liberal (random mixing of piglets in both farrowing and nursery facilities with large pens in the nursery facilities) and the most restrictive strategies (no cross fostering and pigs grouped by litters in small pens in the nursery facilities). These results were compared with actual seroprevalence data obtained from a field study (figure 3). The predicted average seroprevalences from simulations with the liberal strategy were close to those recorded in farms with clinical cases of PMWS, whereas the seroprevalence pattern with the restrictive strategy was similar to that observed in sub-clinically infected farms (figure 3).
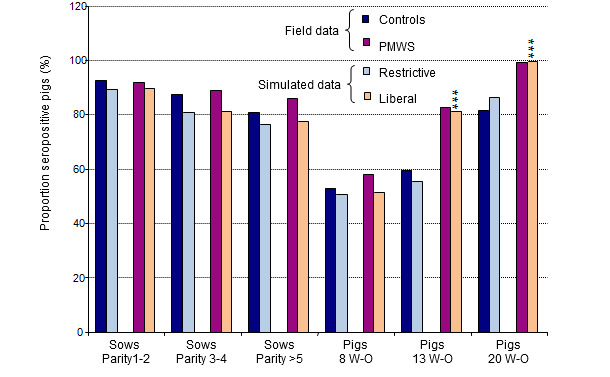
Figure 3. Comparison of outputs (PCV2 seroprevalence) from the simulation model with field data
Evaluating vaccination strategies, the simulation results showed that piglet-targeted vaccines greatly reduced the number of infections. This implies a strong protective effect of piglet vaccination leading to a dramatic decrease in the number of PCV2 infections in the nursery and fattening compartments. Hence, by showing a considerable decrease of the number of early PCV2 infections, these results could explain the reported strong efficacy of PCV2 vaccines on clinical PMWS if the main cause of the disease is controlled to such an extent.
Through this PCV2 example, the modelling approach has shown its powerful contribution to the understanding of a complex disease due to a specific course of infection of the involved pathogen. When properly built, with a careful estimation of parameters, modelling studies are extremely useful to understand biological mechanisms, evaluate scenarios and suggest new research questions.




